Note
Click here to download the full example code
Compare DuBE with resampling-based IL methods (5 classes)
In this example, we compare the duplebalance.DupleBalanceClassifier
and other resampling-based class-imbalanced learning methods.
print(__doc__)
RANDOM_STATE = 42
Preparation
First, we will import necessary packages and generate an example multi-class imbalanced dataset.
from duplebalance import DupleBalanceClassifier
from duplebalance.baselines import ResampleClassifier
from duplebalance.base import sort_dict_by_key
import pandas as pd
from collections import Counter
import matplotlib.pyplot as plt
from sklearn.tree import DecisionTreeClassifier
from sklearn.datasets import make_classification
from sklearn.model_selection import train_test_split
from sklearn.metrics import roc_auc_score
Make a 5-class imbalanced classification task
X, y = make_classification(n_classes=5, class_sep=1, # 5-class
weights=[0.05, 0.05, 0.15, 0.25, 0.5], n_informative=3, n_redundant=1, flip_y=0,
n_features=20, n_clusters_per_class=1, n_samples=2000, random_state=0)
X_train, X_test, y_train, y_test = train_test_split(X, y, test_size=0.5, random_state=42)
origin_distr = sort_dict_by_key(Counter(y_train))
test_distr = sort_dict_by_key(Counter(y_test))
print('Original training dataset shape %s' % origin_distr)
print('Original test dataset shape %s' % test_distr)
Out:
Original training dataset shape {0: 52, 1: 48, 2: 145, 3: 268, 4: 487}
Original test dataset shape {0: 48, 1: 52, 2: 155, 3: 232, 4: 513}
Train DuBE Classifier with different ensemble size
n_estimators_list = [1, 3, 5, 10, 20]
ensemble_init_kwargs = {
'random_state': RANDOM_STATE,
}
eval_kwargs = {'average': 'macro', 'multi_class': 'ovo'}
BASECLF = DecisionTreeClassifier()
# Initialize results list
all_results = []
for n_estimators in n_estimators_list:
# Train DuBE classifiers
clf = DupleBalanceClassifier(
base_estimator=BASECLF,
n_estimators=n_estimators,
**ensemble_init_kwargs
).fit(
X_train, y_train,
resampling_target='under',
resampling_strategy='shem',
perturb_alpha=.5,
sample_weight=None,
eval_datasets={'test': (X_test, y_test)},
train_verbose=False,
)
y_pred_proba = clf.predict_proba(X_test)
score = roc_auc_score(y_test, y_pred_proba, **eval_kwargs)
print ("DuBE {:<2d} | Balanced AUROC: {:.3f} | #Training Samples: {:d}".format(
n_estimators, score, sum(clf.estimators_n_training_samples_)
))
all_results.append(
['DuBE', score, sum(clf.estimators_n_training_samples_)]
)
Out:
DuBE 1 | Balanced AUROC: 0.869 | #Training Samples: 240
DuBE 3 | Balanced AUROC: 0.945 | #Training Samples: 720
DuBE 5 | Balanced AUROC: 0.965 | #Training Samples: 1200
DuBE 10 | Balanced AUROC: 0.977 | #Training Samples: 2400
DuBE 20 | Balanced AUROC: 0.984 | #Training Samples: 4800
Train Resampling-based IL Classifiers
from imblearn.under_sampling import *
from imblearn.over_sampling import *
resamp_init_kwargs = {'random_state': 42}
samplers = {
'No-resampling': None,
'RUS': RandomUnderSampler(**resamp_init_kwargs),
'TomekLinks': TomekLinks(),
'NearMiss': NearMiss(),
'Condense': CondensedNearestNeighbour(**resamp_init_kwargs),
'ROS': RandomOverSampler(**resamp_init_kwargs),
'SMOTE': SMOTE(**resamp_init_kwargs),
'ADASYN': ADASYN(**resamp_init_kwargs),
'BorderSMOTE': BorderlineSMOTE(**resamp_init_kwargs),
}
# Train all resampling-based imbalanced learning methods
for sampler_name, sampler in samplers.items():
clf = ResampleClassifier(
base_estimator=BASECLF,
sampler=sampler
).fit(X_train, y_train)
y_pred_proba = clf.predict_proba(X_test)
score = roc_auc_score(y_test, y_pred_proba, **eval_kwargs)
print (sampler_name, score, clf.n_training_samples_)
all_results.append(
[sampler_name, score, clf.n_training_samples_]
)
Out:
No-resampling 0.8828851091981544 1000
RUS 0.8727953972366624 240
TomekLinks 0.8804842564696351 920
NearMiss 0.8366551576942509 240
Condense 0.8081403968288517 227
ROS 0.8561541415871521 2435
SMOTE 0.8617858533127144 2435
ADASYN 0.8698936401528098 2390
BorderSMOTE 0.8392594246977406 2435
Results Visualization
import matplotlib.pyplot as plt
import seaborn as sns
sns.set_context('talk')
def plot_results_comp(data_vis, x, y, title, figsize=(8,6)):
fig = plt.figure(figsize=figsize)
ax = sns.lineplot(
data=data_vis, x=x, y=y, hue='Method', style='Method',
markers=True, err_style='bars', linewidth=4, markersize=20, alpha=0.9
)
for position, spine in ax.spines.items():
spine.set_color('black')
spine.set_linewidth(2)
ax.grid(color = 'black', linestyle='-.', alpha=0.3)
ax.set_ylabel('AUROC (macro)')
ax.set_title(title)
ax.legend(
title='',
borderpad=0.25,
columnspacing=0.05,
borderaxespad=0.15,
handletextpad=0.05,
labelspacing=0.05,
handlelength=1.2,
)
return ax
all_results_columns = ['Method', 'AUROC (macro)', '#Training Samples']
data_vis = pd.DataFrame(all_results, columns=all_results_columns)
plot_results_comp(data_vis, x='#Training Samples', y='AUROC (macro)',
title='DuBE versus Resampling Baselines')
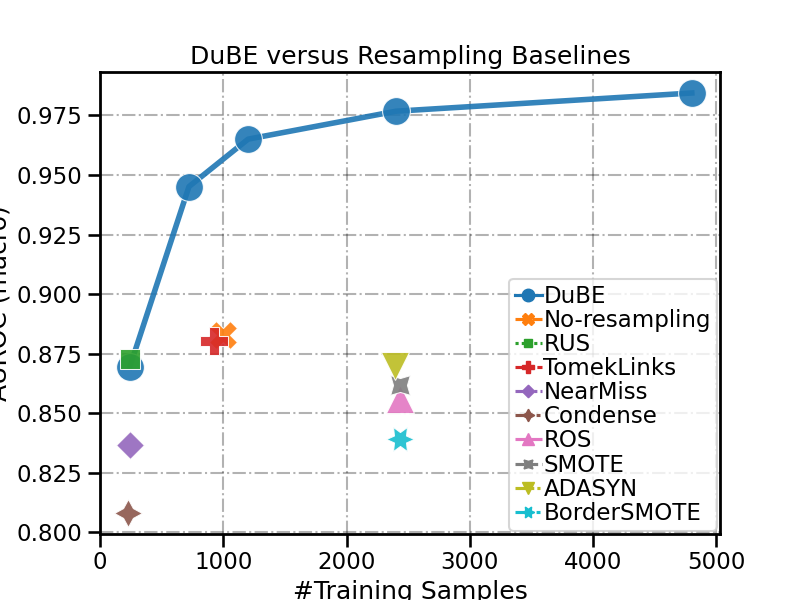
Out:
<AxesSubplot:title={'center':'DuBE versus Resampling Baselines'}, xlabel='#Training Samples', ylabel='AUROC (macro)'>
Total running time of the script: ( 0 minutes 44.053 seconds)
Estimated memory usage: 15 MB